Variant QC and statistics¶
The Variant QC and statistics report builder generates a quality control report for sequence data in selected subjects. Quality parameters returned include coverage per variant, variant genotype, and allele frequencies across subjects, as well as Hardy Weinberg Equilibrium p-values and log p-values per variant.
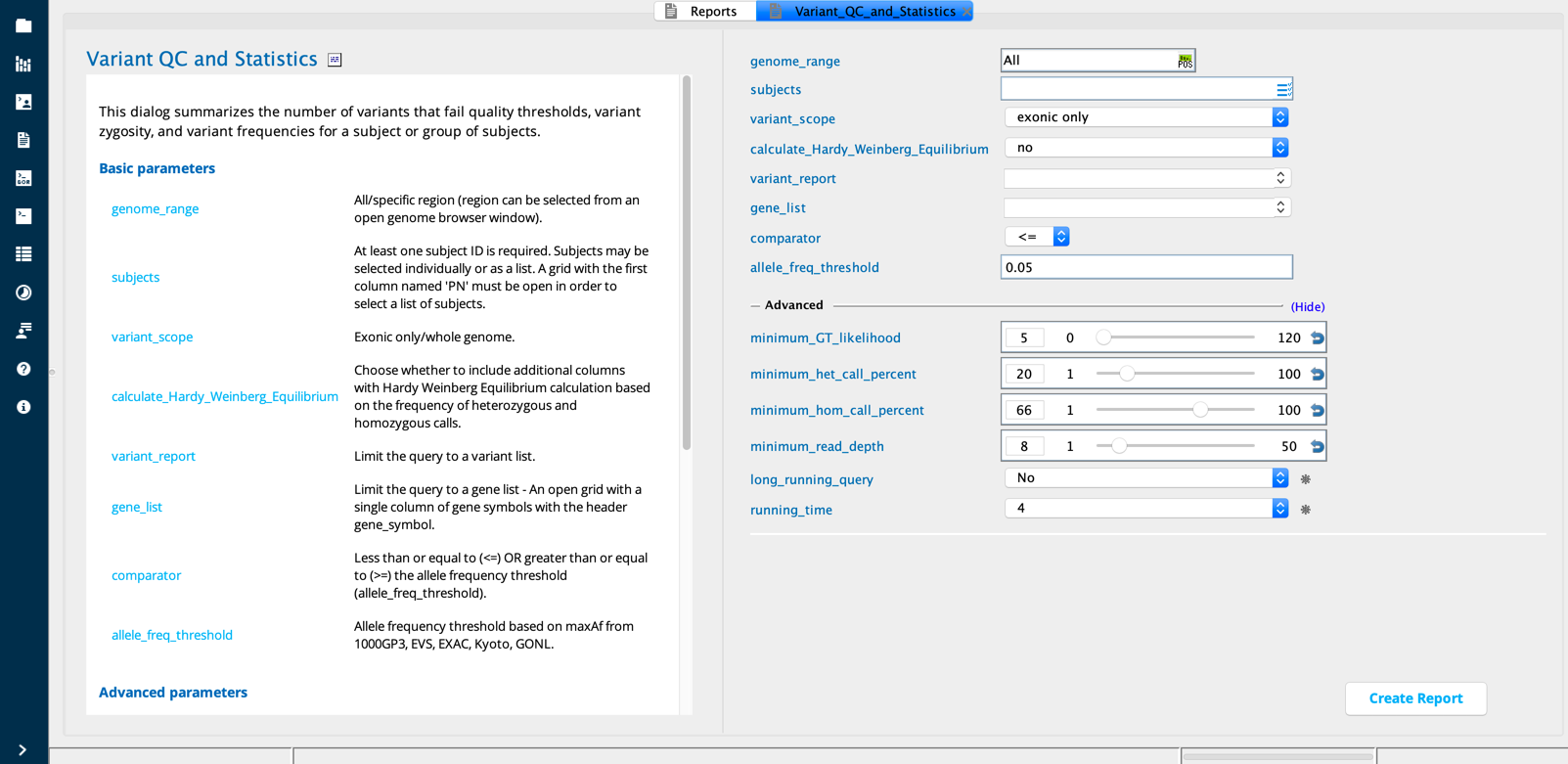
Variant QC and Statistics module in Sequence Miner¶
Example use case¶
The user has a cohort of 300 cases and controls and wishes to evaluate overall data quality and to identify variants not in HWE.
Interpreting the output¶
Given a list of subjects, this dialog summarizes the following for each variant in these subjects:
The number of subjects failing QC at this locus (FailQc_All)
The number of subjects failing QC but having depth of coverage greater than the user-defined threshold at this variant locus (FailQc_NonDepth)
The sum of hom subjects for this variant
The sum of het subjects for this variant
- PNcount - The greater number of subjects calculated under the following two conditions:
The number of subjects carrying this variant with GL likelihood score and call ratio meeting the user-defined thresholds
The number of subjects for which the variant was called as het or hom, which tests GL likelihood score, call ratio, AND read depth
If the user chooses the option to include a Hardy Weinberg Equilibrium calculation, then the following columns are included in the output:
Chi-square value
Hardy Weinberg Equilibrium p-val
Hardy Weinberg Equilibrium log p-val
The frequency of hom subjects
The frequency of het subjects
The frequency of variant alleles among all alleles in the subjects
The sum_het and sum_hom columns report the number of heterozygous and homozygous carriers among the input subjects.
QC is categorized as failing for a given variant in a subject when any of the following parameters fail to meet the user-defined thresholds:
variant GL score (genotype likelihood score),
minimum read depth and
call ratios for the het or hom call
The sum_FailQc_All column contains the number of selected subjects for which the variant fails the user-defined QC (quality) thresholds.
The sum_FailQc_NonDepth column reports the number of subjects that fail QC due to reasons other than read depth being less than the user-defined read depth threshold.
If “yes” is selected in the calculate_Hardy_Weinberg_Equilibrium (calcHWE) field, then the output includes several additional columns. The chi-square test statistic (chisq column) is calculated based on the heterozygous frequency, homozygous frequency, and total allele frequency with one degree of freedom. The corresponding chi-square p Value (pVal) is returned along with the -log(p-value). The -log(p value) can then be plotted in the Genome Browser as a Manhattan plot across the genome.
Note
Expected counts for the HWE calculation are determined from the allele frequencies in the input subjects. Therefore, it is recommended that the HWE calculation be selected only in the case of a large number of samples. Otherwise, Fisher’s exact test is the recommended method for measuring the distribution of the heterozygotes and homozygotes.
Column descriptions¶
Group |
Column |
Description |
|---|---|---|
Basic |
Call |
|
Chrom |
||
POS |
||
Reference |
||
sum |
FailQc_All |
The number of samples that do not pass any of the user-defined thresholds for QC at this variant locus |
FailQc_NonDepth |
The number of samples that do not pass the user-defined thresholds for QC other than read depth at this variant locus |
|
het |
The number of samples heterozygous for the variant |
|
hom |
The number of samples homozygous for the variant |
|
Other columns |
max_af |
|
PNcount |
The number of samples with good coverage (depth meeting the user-defined threshold) at this locus; if variant is present, the variant meets the user-defined thresholds for variant GL score (genotype likelihood score) and call ratios for the het or hom calls |
Additional columns¶
If the calculate_Hardy_Weinberg_Equilibrium (calcHWE) option is set to “yes”, the following columns are added:
Group |
Column |
Description |
|---|---|---|
HWE |
alleleFreq |
(2 * sum_hom + sum_het) * 0.5 / (total number of samples containing this variant), the frequency of the variant |
chisq |
Chi-square test statistic for Hardy Weinberg equilibrium calculated based on the homFreq and homFreq |
|
homFreq |
sum_hom / (total number of samples), the ratio of the number of samples containing this variant in a homozygous state versus the total number of samples |
|
logPval |
Calculated based on the pVal, -log(pVal) |
|
pVal |
The pVal for Hardy Weinberg calculated based on chi-square test statistic (chisq) and 1 degree of freedom |
|
Other columns |
hetFreq |
sum_het / (total number of samples ), the ratio of the number of samples containing this variant in a heterozygous state versus the the total number of samples |