Sample QC and statistics¶
The Sample QC and statistics report calculates variation statistics for autosomal chromosomes and gene statistics for autosomes and sex chromosomes. The QC statistics are reported for selected samples as compared to the distribution of the QC attribute values for all samples in the project.
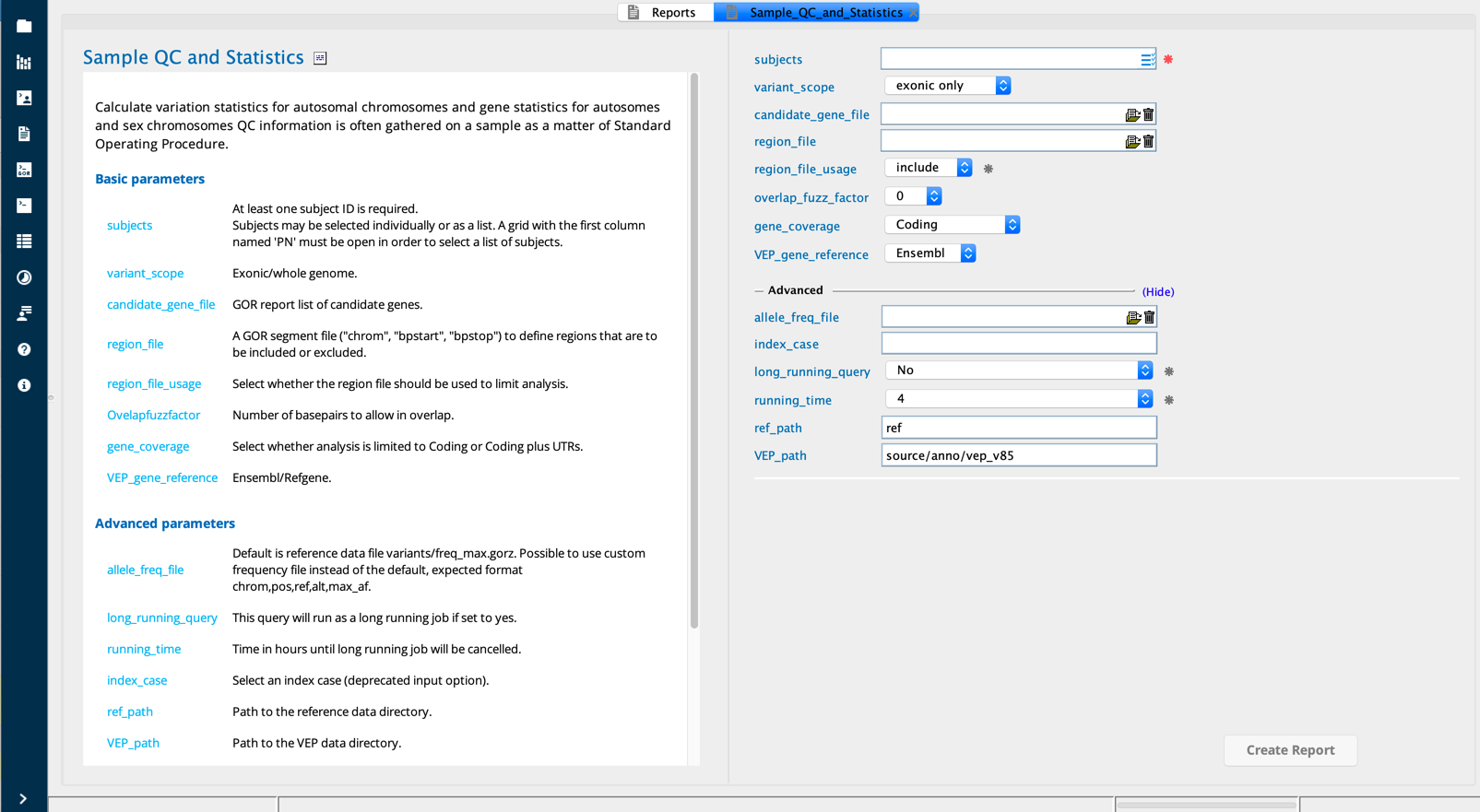
Sample QC and Statistics module in Sequence Miner¶
Example use case¶
The user wishes to generate a detailed report of variant and gene QC statistics for a list of selected subjects.
Description of the algorithm¶
For each subject loaded into the WuXi NextCODE platform, several files are generated, including BAM files, VCF files, per-base coverage files, a good coverage locus file, and VEP scores for variant. This report builder draws from all of these files to generate a detailed QC report.
In addition to the Default view perspective, there are five other Perspective views focusing on exome and gene coverage, and on a breakdown of variant attributes per sample. Frequencies, distributions, and additional statistical analyses are generated for attributes.
Interpreting the output¶
The Default view perspective reports all QC statistics and displays results for nine analyses:
SNP vs InDel Analysis: The fraction of variants identified as “SNP” or “InDel”; the proportions of SNP and InDels are expected to be >.96 and > 0.03, respectively
dbSNP Analysis: The fraction of variants that were found in dbSNP; the typical proportion of “absent” and “present” dbSNP variants is expected to be < 0.1 and 99.9, respectively.
exomeCoverage: Coverage for exons in the exome of a given sample is determined by mapping the transcript coordinates from the reference gene set (Ensembl/Refgene) mapped to a per base coverage file. The coverage across the exome is reported as a value between 0 and 1 that indicates the proportion of the exome that has less than the given coverage value (e.g., exlt10 = fraction with less than 10X coverage). This is intended to flag samples with a higher than usual proportion of the exome with low coverage (e.g., a value of 0.8 for exlt10 indicates 80% of the exome for that sample has less than 10X coverage).
frequency Analysis: The fraction of variants found in each of the following categories: “veryrare” (AF < 0.1%), rare (AF 0.1% - 1%), “medium” (AF 1% - 5%), and “common” (AF > 5%)
geneCoverage: Coverage for the gene is determined based on mapping of the per base coverage file for each sample to the coordinates of the genes in the selected reference set (Ensembl or Refgene). The proportion of all genes for a given sample that contain a minimum coverage is reported as a value between 0 and 1. This attribute is determined based on what percentage of the gene has a minimum level of coverage (either 10X or 20X) and is reported as the proportion of all genes having 85%, 90%, or 95% of a gene covered by 10X or 20X.
- impact Analysis: The fraction of variants found in each of the following VEP impact categories:
HIGH impact (typically expected to contain approximately < 0.02 of variants)
MODERATE impact (typically expected to contain approximately 0.40-0.45 of variants)
LOW impact (typically expected to contain approximately 0.40-0.45 of variants)
LOWEST impact (typically expected to contain approximately 0.10-0.15 of variants)
quality Analysis: Variant call quality, either “LowQual” or “PASS”; the proportions of LowQual and PASS variants are expected to be < 0.1 and 99.9, respectively
transition transversion Analysis: A transition refers to a purine to purine substitution (A->G) or pyrimidine to pyrimidine substitution (C->T). A transversion refers to a purine to pyrimidine substitution (A->C, A->T, G->C, or G->T) or vice versa. A transition to transverstion calculation provides a readout of the total number of transitions called versus the total number of transversions called.
zygosity Analysis: The fraction of variants that are heterozygous (het) and homozygous (hom); expected to be approximately 0.6 and 0.4, respectively
The attribute measurement for each subject is compared to its distribution across the other input SUBJECTS. The distribution is expressed as a z-score, the number of standard deviations from the mean value for a given attribute. The Z-score value is displayed in the all_z_proportion column. The corresponding rank (LOW, NORMAL and HIGH) of the attribute value for each SUBJECT compared to the distribution of all the input SUBJECTS is displayed in the all_InDistribution column. The all_InDistribution column indicates how a sample attribute (in a given row) compares to the other samples (SUBJECTS) selected for this analysis.
If the Z-score (the value in the all_z_proportion column is the number of standard deviations away from the mean) is < -2.0 (more than 2 standard deviations BELOW the mean), the attribute is categorized as falling at the LOW end of the distribution of all the input SUBJECTS (designated in the all_InDistribution column).
If the Z-score is > +2.0 (more than 2 standard deviations ABOVE the mean), the attribute is categorized as falling at the HIGH end of the distribution of all the input SUBJECTS.
If the Z-score is between -2.0 and and +2.0 (within 2 standard deviations of the mean), the attribute is categorized falling within the NORMAL range of the distribution of all the input SUBJECTS.
Column descriptions¶
Group |
Column name |
Description |
|---|---|---|
Basic |
Chrom |
|
PN |
||
All |
avg_avg_depth |
Across all samples in the analysis, the average of the average of the 85, 90 and 95% gene depth coverage at greater than 10 and 20 reads |
avg_depth |
Per sample, the average of 85, 90 and 95% gene depth coverage at greater than 10 and 20 reads |
|
avg_proportion |
For each gene depth measurement category, the average value across all selected SUBJECTS is calculated (only visible in the GeneStatGraphs perspective) |
|
Color |
|
|
exomeSize |
Total exome size of the sample (reported in base pairs) |
|
InDistribution |
|
|
lowOReqRankFromTop |
Samples are ranked proportionally on a scale of 0 to 1 (each step in rank is 1 / # of samples), with 0 being the highest rank assigned to the sample with the highest attribute value |
|
lowOReqRankFromBottom |
Samples are ranked proportionally on a scale of 0 to 1, with 0 being the highest rank assigned to the sample with the lowest attribute value |
|
numberOfGenes |
Total number of genes found in the sample for the “All” or “Candidate” genes designation |
|
PNcount |
The total count of samples in the analysis |
|
proportion |
Percentage of total variations having the designated value in the attribute column for the analysis performed (value between 0 and 100) |
|
rank_perc_FromTop |
Samples are ranked in order based on their attribute values (e.g., 1 - 21 for 21 samples), with 1 being the highest rank assigned to the sample with the highest attribute value |
|
rank_perc_FromBottom |
Samples are ranked in order based on their attribute values, with 1 being the highest rank assigned to the sample with the lowest attribute value |
|
ratio |
The ratio of the indicated numerator (e.g., SNPs, transitions, heterozygous) to the denominator (e.g., InDels, transversions, homozygous) |
|
std_avg_depth |
Standard deviation of the average coverage at all exome and gene coverage values across all SUBJECTS selected for the analysis |
|
std_proportion |
For each gene depth measurement category, the standard deviation of the average value across all selected SUBJECTS is calculated (only visible in the GeneStatGraphs perspective) |
|
totalVars |
Total variants contributing to the ratio calculation |
|
z_proportion |
Z-score value |
|
Other columns |
analysis |
|
Attribute |
||
bpStart |
Perspective views¶
In addition to the Default view perspective, results are displayed in five other perspectives focusing on exome and gene coverage, and on a breakdown of variant attributes per sample. Frequencies, distributions, and additional statistical analyses are generated for attributes.
Perspective |
Description |
|---|---|
Default view |
Reports all QC statistics and displays results for the 9 anaylses described above |
ExomeCovTable |
Reports the coverage statistics for exons |
GeneStatGraphs |
Reports gene coverage statistics |
GeneStatTable |
Reports gene coverage statistics |
Ratios |
Includes 3 categories of ratios listed in the analysis column (other analyses are filtered out): zygosity Analysis, SNP vs InDel Analysis, and transition transversion Analysis |
VariantStat |
Includes 6 categories of variant statistics listed in the analysis column: zygosity Analysis, SNP vs InDel Analysis, dbSNP Analysis, frequency Analysis, impact Analysis, and quality Analysis |