Risk SNPs¶
The Risk SNPs report builder annotates the variants from a PN or list of PNs that match a set of user-defined rsIDs from the dbSNP reference file.
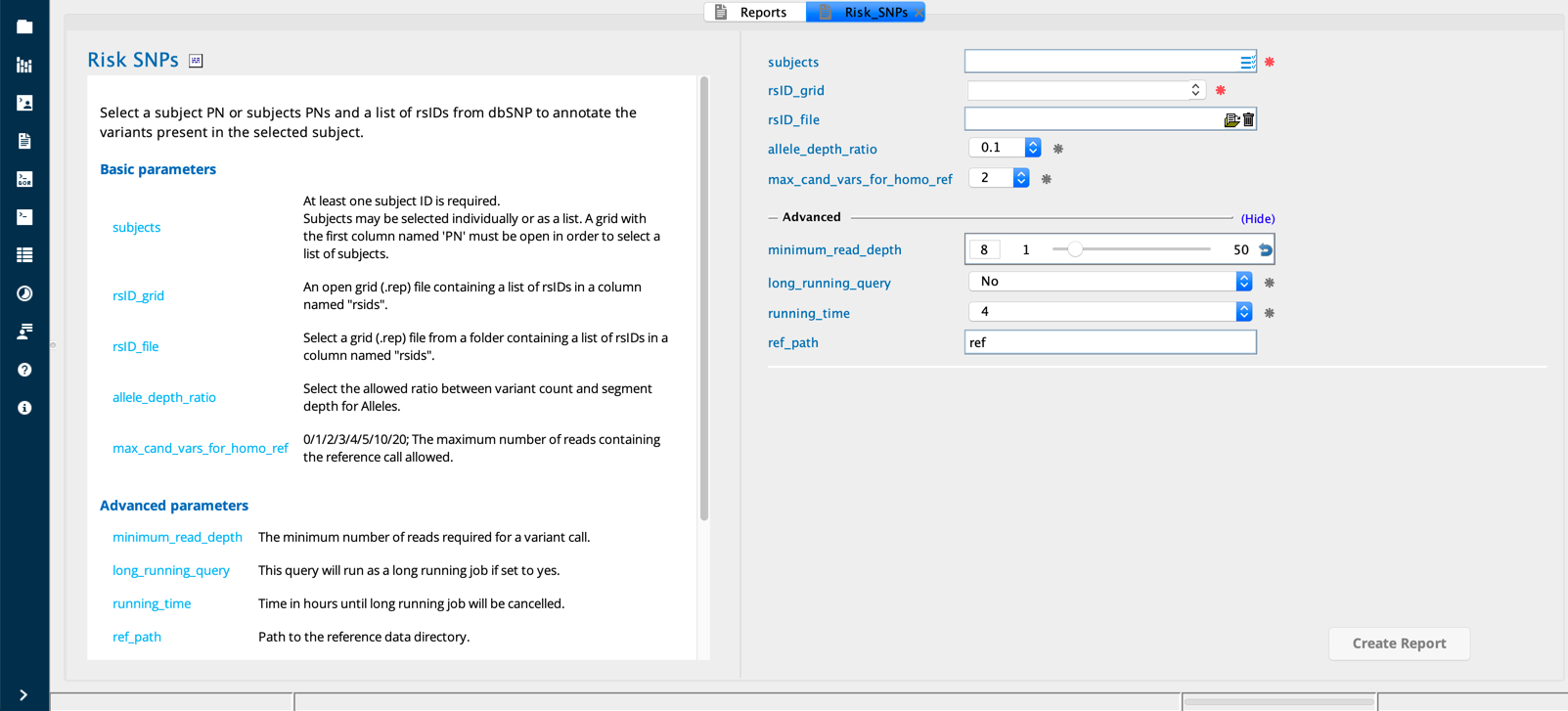
Risk SNPs module in Sequence Miner¶
Example use case¶
The user has a list of rsIDs for SNPs of interest (from dbSNP) and wishes to screen the subject(s) for those variants and to determine the quality of the variant if present.
Description of the algorithm¶
rsIDs matching the user-designated rsID_grid are extracted from the dbSNP reference file (ref/dbsnp/dbsnp_anno.gorz). This table is expanded by creating one row per subject (PN) per rsID.
Each rsID/SNP coordinate for each subject is then joined with the corresponding coordinate and PN (row) in the source/var/wgs_varcalls.gord file.
Allele calls (e.g., “A/T”) for each rsID for each PN are reported in the Alleles column if either of the following sets of conditions is true. Otherwise, the call/call is changed to “Unknown”.
Condition set 1
The variant call is listed as “Missing”
Read depth is > the user designated minReadDepth
There are < 2 reads for the variant OR (reads for the variant) / (reads for the segment ) < 0.10
Condition set 2
The variant call is NOT listed as “Missing”
Read depth is > the user designated minReadDepth
Interpreting the output¶
The resulting table provides the following information:
If the selected sample carries the variant
Read depth of the variant
Read depth of that locus
Raw reads containing the variant
The readsWithNonRef columns display the read depth of the allele derived from the VCF - “0” indicates that this variant is not carried by the selected subject.
The Depth columns display the total read depth of the locus in that particular sample, which is the DP value in the VCF file - “unknown” indicates that this variant does not exist in that particular sample.
The values displayed in the Depth and readsWithNonRef columns are different:
The readsWithNonRef column is calculated from the raw read count file named
candidate_variants.gordand contains a pile-up of the number of reads containing the variant allele in the BAM.The Depth column is the total read depth for the given position and is derived from the VCF file DP column (
source/var/wes_varcalls.gord).
The SegDepth column displays the raw read depth of the locus calculated from the BAM in the per base coverage file pileup, which is extracted from the segment_cov.gord file. This value can be used to determine if there is sufficient read coverage of the locus to validate the variant call.
Column descriptions¶
Group |
Column |
Description |
|---|---|---|
Basic |
Chrom |
The chromosome of the variant represented as chr1, chr2, …, chr22, chrXY, chrX, chrY, chrM |
PN |
The patient number (identifier) |
|
POS |
The (first) base pair position of the sequence variant, i.e., the position of the first nucleotide in the Reference column |
|
Other columns |
Alleles |
“unknown” indicates that this rsID-related variant is not present in the selected sample because the coverage is too low or there are insufficient reads containing the variant |
Call |
“missing” indicates that this rsID-related variant is not present in the selected sample |
|
dbSNP_Alleles |
the matching variant alleles from the dbSNP reference database |
|
Depth |
The total read depth reported in the VCF file |
|
readsWithNonRef |
The number of reads that contain the variant, calculated based on the candidate_variants.gord file (based on BAM file) |
|
rsID |
The rsID from the input file |
|
SegDepth |
The total read depth reads that cover the locus, which is calculated based on segment_cov.gord file (based on BAM file) |
|
Strand |
Chromosome strand, either “+” or “-” to indicate the direction in which the variant sequence is transcribed |
Perspective views¶
Perspectives subtabs focus on subsets of the columns in the Default view.
Perspective |
Description |
|---|---|
Basic |
|
Default view |
Shows all columns |