Multi-family Mendelian analysis¶
The Multi-family Mendelian analysis report builder returns the zygosity distribution across cases and controls (the number of homozygous, heterozygous, and compound heterozygous carriers) for each variant observed in the index case.
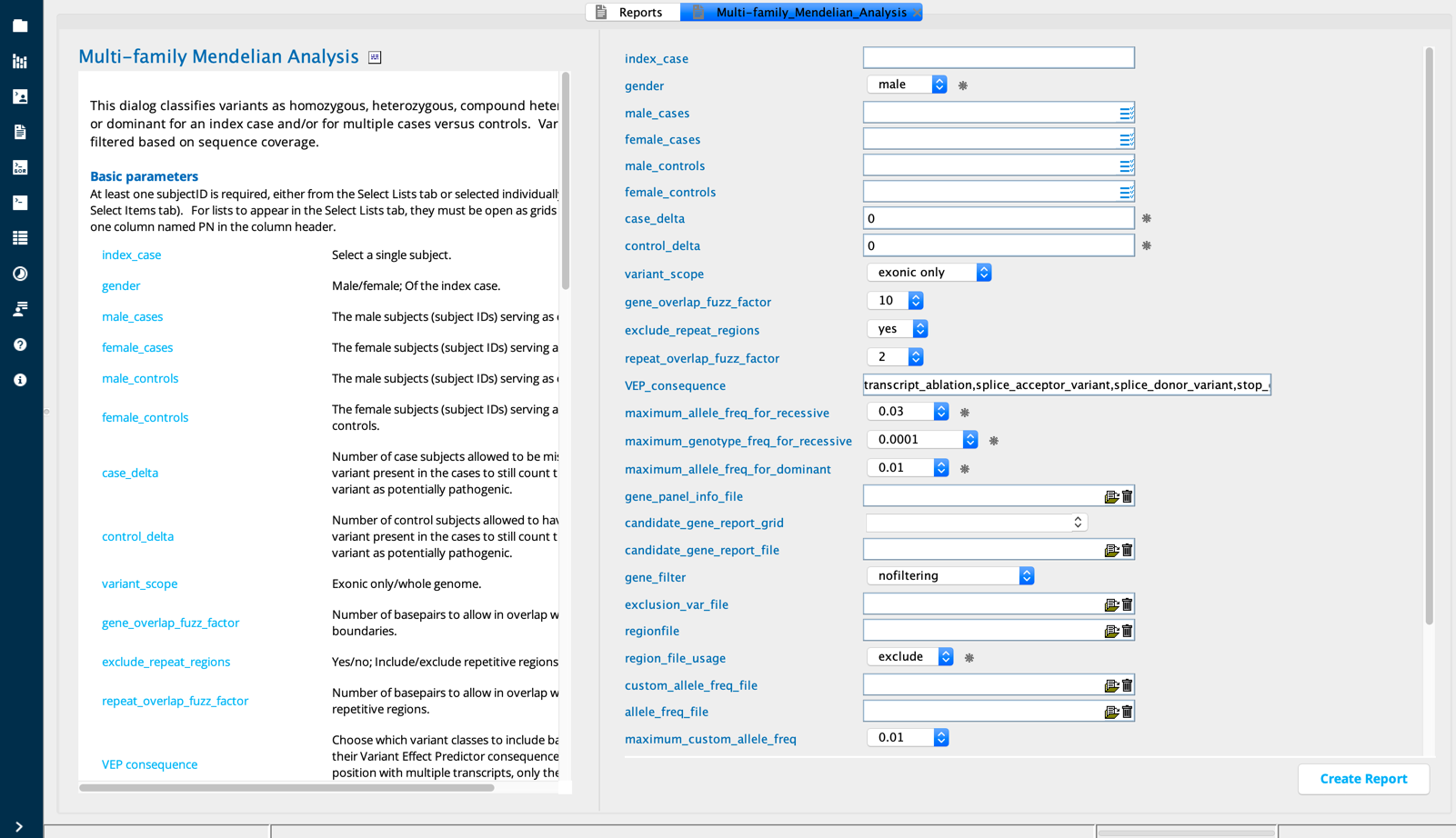
Multi-Family Mendelian Analysis in Sequence Miner¶
Example use case¶
Given an index case and a group of cases and controls, the user wishes to evaluate the variants observed in the index by measuring the distribution of homozygous, heterozygous, and compound heterozygous carriers of each variant across cases compared to controls.
Description of the algorithm¶
Variants are first filtered according to coverage and other user-defined settings. Variants are classified as homozygous, heterozygous or compound heterozygous for an index case and/or for multiple cases versus controls.
Interpreting the output¶
The output is a detailed report with annotation for every variant identified in the index case that meets the filtering criteria selected by the user (VEP_consequence, maxAf cutoffs for dominant and recessive models, maxGf for the recessive model, etc.). Annotation includes sample and data quality metrics, VEP, and diagnostic information.
Column descriptions¶
Group |
Column |
Description |
|---|---|---|
Basic |
||
Call |
The actual called sequence (variant), found by replacing a part of the reference sequence and denoted by Pos and Reference, with the sequence in the Call column |
|
Chrom |
he chromosome of the variant, represented as chr1, chr2, …, chr22, chrXY, chrX, chrY, chrM |
|
hetORhom |
The zygosity of the call, either “het” or “hom” |
|
Pos |
The (first) base pair position of the sequence variant, i.e., the position of the first nucleotide in the Reference column |
|
Reference |
Sequence from the reference build; the first base starting at the base pair position in the Pos column |
Group |
Column |
Description |
|---|---|---|
CGD |
The CGD columns provide information for variants based on the manually curated database of variants associated with known medically significant conditions and available interventions (http://research.nhgri.nih.gov/CGD/) |
|
AGE_GROUP |
Pediatric: less than 18 years of age; Adult: at least 18 years of age |
|
COMMENTS |
Observations noted by curators |
|
CONDITION |
Conditions resulting from mutations in the same gene but may otherwise be placed in the “General” Intervention category |
|
INHERITANCE |
Pattern of inheritance: AD - autosomal dominant; AR - autosomal recessive; BG - blood group; Digenic - a condition resulting from simultaneous mutations in two different genes; Maternal - maternal mitochondrial inheritance; XL - X-linked (because X-linked conditions can frequently have manifestations in both genetic sexes, X-linked conditions are not designated as dominant or recessive) |
|
INTERVENTION_CATEGORIES |
This category includes organ systems for which specific and additional interventions may be beneficial |
|
INTERVENTION_RATIONALE |
Description of the intervention and its benefit |
|
MANIFESTATION_CATEGORIES |
Includes organ systems affected by mutations in corresponding genes; recognition of involved organ systems may help guide supportive care |
|
REFERENCES |
The PubMed ID (PMID) of the reference |
Group |
Column |
Description |
|---|---|---|
Diag |
||
ACMGcat |
Categorization of the sequence variants according to the ACMG scheme |
|
chz |
This field is equal to 1 (true) if the index is/cases are compound heterozygous (CHZ) (see GENE_CHZinGene) and none of the controls are homozygous |
|
ChzAHet |
CHZ with allelic heterogeneity (AHet); this field is equal to 1 (true) if the variant contributes to CHZ in a gene (not necessarily all the same variants) and is absent as a CHZ variant from controls (in males, the X chromosome is considered separately) |
|
Dom |
Dominant variant; this field is equal to 1 (true) if the variant is present in cases and absent in control individuals |
|
DomAhet |
Dominant variant considering allelic heterogeneity (AHet); this field is equal to 1 (true ) if variants in a gene (not necessarily the same variant) are present in cases (“f/m_subjWithVarInGene” > 1) and absent in control subjects (“f/m_CTRLs_subjWithVarInGene” = 0) |
|
HRec |
Homozygous recessive variant; this field is equal to 1 (true) if the variant is homozygous in cases (or hemizygous in males for variants on the X chromosome) and no homozygous variants (or hemizygous variants on X in males) are present in controls |
|
HRecAHet |
Homozygous recessive with allelic heterogeneity (AHet); this field is equal to 1 (true) if the variant is homozygous in cases, or males are hemizygous for variants on the X chromosome (not necessarily the same variant in all cases) and no homozygous variants (or hemizygous variants on X in males) are present in controls |
|
otherpos |
For a particular CHZ variant, the basepair position of the other variant(s) that produces compound heterozygosity |
Group |
Column |
Description |
|---|---|---|
EuroGenetest |
The EuroGenetest columns are derived from a European Commission project database containing European genetic testing information for particular genes, variants, and diseases. |
|
Diseases |
Diseases associated with a variant derived from the European Commission project database |
|
NoOfDiseases |
Number of diseases associated with a variant derived from the European Commission project database |
|
NoOfpanels |
Number of gene panels associated with a variant derived from the European Commission project database |
|
panels |
EuroGenetest panels associated with a variant derived from the European Commission project database |
Group |
Column |
Description |
|---|---|---|
female |
In the female group, the following columns are repeated for fCASEs and fCTRLs. |
|
GeneCovered |
The number of subjects (within the given male/female and case/control category) that have at least 8 reads (depth >=8) at the position of all the variants in a given gene |
|
subjCompHeterInGene |
The number of subjects (within the given male/female and case/control category) who are potentially compound heterozygous in a given gene. Note that the variants are not phased based on parent of origin; an individual need only have a single homozygous variant or two heterozygous variants in a given gene to be classified as CHZ |
|
subjWithHomVar: |
The number of subjects (within the given male/female and case/control category) homozygous for the given variant |
|
subjWithHomVarInGene |
The number of subjects (within the given male/female and case/control category) with homozygous variants in the gene |
|
subjWithVar |
The number of subjects (within the given male/female and case/control category) with the variant |
|
subjWithVarInGene |
The number of subjects (within the given male/female and case/control category) with any variant in the gene |
|
varCovered |
The number of subjects (within the given male/female and case/control category) that have at least 8 reads at the variant position |
Group |
Column |
Description |
|---|---|---|
Gene |
||
Aliases |
The aliases of the given gene |
|
avg_depth |
The average sequence read depth in the exome of the given gene |
|
exomeSize |
The sum of the base pair size of all the exons in the gene |
|
exontype |
The type of exon (“coding” or “noncoding”) |
|
lt10 |
The fraction of the exome with sequence read coverage less than 10X |
|
lt15 |
The fraction of the exome with sequence read coverage less than 15X |
|
lt20 |
The fraction of the exome with sequence read coverage less than 20X |
|
lt25 |
The fraction of the exome with sequence read coverage less than 25X |
|
lt30 |
The fraction of the exome with sequence read coverage less than 30X |
|
lt5 |
The fraction of the exome with sequence read coverage less than 5X |
|
maximum_allele_freq_for_dominant |
||
maximum_allele_freq_for_recessive |
||
maximum_genotype_freq_for_recessive |
||
MOI |
Mode of inheritance |
|
Paralogs |
The paralogs of the given gene |
|
RmaxAf |
Maximum allele frequency to use for recessive alleles set as input parameter |
|
RmaxGf |
Maximum gene frequency to use for recessive disorders set as input parameter |
|
Symbol |
Based on HGNC when it exists, otherwise it is the Ensembl internal alias |
Group |
Column |
Description |
|---|---|---|
GO |
||
Descriptions |
Gene ontology descriptions |
|
IDs |
Gene ontology identifiers |
Group |
Column |
Description |
|---|---|---|
GT |
||
CallCopies |
Because the focus is only on variants from the reference, CallCopies refers to how many copies of the variant exist in a subject; a CallCopies value of 2 therefore corresponds to a homozygous variant, whereas a CallCopies value of 1 corresponds to a heterozygous variant |
|
CallRatio |
Proportion of reads containing the variant call; expected to be close to 0.5 for heterozygous calls and close to 1 for homozygous calls |
|
Depth |
The number of reads used for evaluating the corresponding call |
|
FILTER |
Quality parameter using the ratio between gt-quality and depth, showing if the call is considered “LowQual” quality (not useable) or “PASS”; this remains a crude quality measure |
|
formatZip |
VCF genotype field |
|
FS |
Fisher’s exact test of read strand; f the reference reads are balanced between forward and reverse strands, then the alternate reads should be as well |
|
GL_Call |
A statistical measure indicating the likelihood that the call is wrong; the scale has been converted to use only integer numbers - the higher the number, the less likely it is that the call is wrong |
|
vcf_alt |
Alternate (variant) nucleotide sequence as reported in the vcf |
|
vcf_pos |
Coordinate (position) of the reference nucleotide sequence |
Group |
Column |
Description |
|---|---|---|
KNOWN |
The KNOWN columns provide publicly available information about the candidate gene and/or variant. |
|
distance |
The distance between a known variant and the identified variant |
|
exactMatch |
A Boolean column (1/0) indicating if the variant is a known variant (instead of near a known variant, or a at the same position with a different call allele) |
|
Gene_diseases |
Diseases known to be associated with the gene |
|
Gene_Symbol |
Based on HGNC when it exists, otherwise it is the Ensembl internal alias |
|
GeneLists |
Gene list membership of the gene in which the variant is found |
|
pmid |
PubMed ID of the reference from which the information was obtained |
|
var_diseases |
Diseases known to be associated with the variant |
Group |
Column |
Description |
|---|---|---|
male |
In the male group, the following columns are repeated for mCASEs and mCTRLs. |
|
GeneCovered |
The number of subjects (within the given male/female and case/control category) that have at least 8 reads (depth >=8) at the position of all the variants in a given gene |
|
subjCompHeterInGene |
The number of subjects (within the given male/female and case/control category) who are potentially compound heterozygous in a given gene. Note that the variants are not phased based on parent of origin; an individual need only have a single homozygous variant or two heterozygous variants in a given gene to be classified as CHZ |
|
subjWithHomVar |
The number of subjects (within the given male/female and case/control category) homozygous for the given variant |
|
subjWithHomVarInGene |
The number of subjects (within the given male/female and case/control category) with homozygous variants in the gene |
|
subjWithVar |
The number of subjects (within the given male/female and case/control category) with the variant |
|
subjWithVarInGene |
The number of subjects (within the given male/female and case/control category) with any variant in the gene |
|
varCovered |
The number of subjects (within the given male/female and case/control category) that have at least 8 reads at the variant position |
Group |
Column |
Description |
|---|---|---|
OMIM |
The OMIM columns provide the OMIM-designated identification for a particular gene and related disease description |
|
Descriptions |
OMIM disease descriptions for the gene |
|
IDs |
The OMIM ID(s) of the gene |
Group |
Column |
Description |
|---|---|---|
VEP |
The VEP (Variant Effect Predictor) columns provide functional annotations for variants based on the ENSEMBL SNP Effect Predictor database. For more information, visit the VEP web page: http://www.ensembl.org/info/docs/tools/vep/index.html/. |
|
CDS_position |
Position of the base pair in the coding sequence; a value is given for each transcript |
|
Max_Af |
Maximum allele frequency from public databases (1000Genomes, Exome Variant server, etc.) |
|
max_consequence |
Consequence type reported for this variant having the greatest impact |
|
Max_Impact |
Classification of the level of severity of the transcript consequence type assigned by VEP |
|
Max_Score |
Maximum score for the variant as observed in dbNSFP [Score=max ((1-Sift_score), Polyphen2_HDIV_score, Polyphen2_HVAR_score)] |
|
Protein_Position |
Position of the amino acid in the protein sequence (only if the variant falls within a coding sequence); a value is given for each corresponding transcript specified in the CDS position field |
|
Transcript_count |
Number of different transcripts in which the variant is found |
Group |
Column |
Description |
|---|---|---|
Other columns |
||
Amino_Acids |
The amino acid with and without variant (provided only if the variation affects the protein-coding sequence), otherwise “.” |
|
Biotype |
Biological class of transcript or regulatory feature |
|
CLINICAL_SIGNIFICANCE |
The clinical significance (e.g., pathogenic, benign, unknown significance, drug-response, risk factor, etc.) of the variant as annotated (commented) by users. If the same variant has several comments, this cell will contain a set of values |
|
MODE_OF_INHERITENCE |
The user-annotated (commented) mode of inheritance of the variant; if the same variant has several comments, this cell will contain a set of values |
|
Refgene |
The accession number from NCBI of the affected transcripts |
|
TEXT |
The description (comment) component for the user’s annotation of the variant |
Perspective views¶
Perspectives subtabs focus on subsets of the columns in the Default view.
Perspective |
Description |
|---|---|
Default view |
|
Dominant |
|
Recessive |
Drill-in reports¶
Drill in |
Description |
|---|---|
CHZforVar |
This drill-in report lists all compound heterozygous variants that include the selected gene variant |
CHZforGene |
This drill-in report lists all compound heterozygous variants that reside in the same gene as the selected gene variant |
AllWithVar |
This drill-in report lists all carriers among the cases and controls of the selected variant |
AllVarsInGene |
This drill-in report lists all carriers among the cases and controls of the selected variant or of variants in the same gene as the selected variant |