Genes to pathways¶
The Genes to pathways report builder matches each gene in a list to a pathway(s) that contains the gene and to other genes in the same pathway(s).
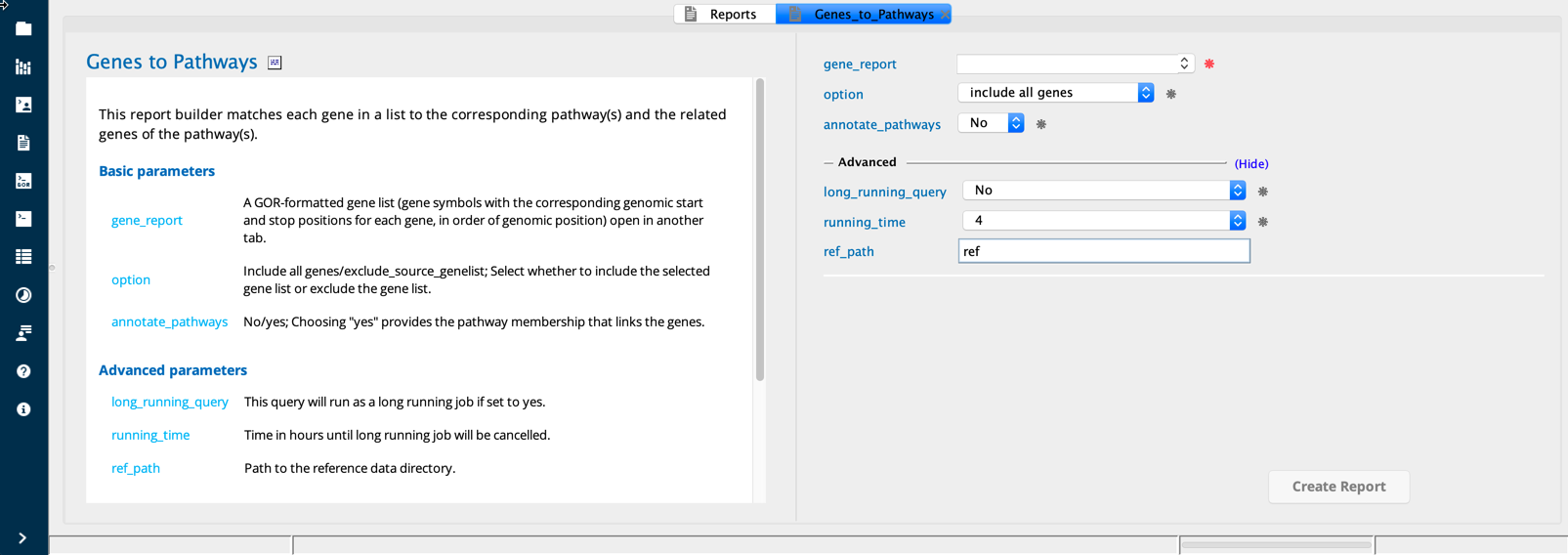
Genes To Pathways module in Sequence Miner¶
Example use case¶
Given a list of genes, the user wishes to determine which pathways contain the genes and all other genes in those pathways.
Description of the algorithm¶
This query takes as input a GOR-formatted genelist that is mapped to a list of pathways containing Ensembl genes. The output lists each gene in the original genelist (if selected for inclusion) with its corresponding pathway (if selected) and a comma-separated list of genes in that pathway.
Interpreting the output¶
Selecting “no” in the annotatePathways field adds a set_orig_gene column in the output, listing genes in the same pathways as the input genelist.
Selecting “yes” in the annotatePathways field adds an additional column, the Pathway column, with the corresponding pathway IDs. Pathway database sources include:
BIOCYC: BioCyc is a collection of 9387 Pathway/Genome Databases (PGDBs).
HGMD: Qiagen’s knowledge base for their Ingenuity Pathway Analysis platform.
KEGG: KEGG PATHWAY is a collection of manually drawn pathway maps representing current knowledge on molecular interaction and reaction networks. KEGG PATHWAY mapping is the process of mapping molecular datasets (e.g., genomics, transcriptomics, proteomics, and metabolomics) to KEGG pathway maps for biological interpretation.
Pathway Interaction Database: The PID is comprised of NCI/Nature-curated pathway information, now available through ndexbio.org hosted by the Ideker Lab at the UC San Diego School of Medicine.
REACTOME: REACTOME is an open-source, open access, manually curated and peer-reviewed pathway database.
WikiPathways: WikiPathways was established to facilitate the contribution and maintenance of pathway information by the biology community. WikiPathways is an open, collaborative platform dedicated to the curation of biological pathways.
Column descriptions¶
Grup |
Column |
Description |
|---|---|---|
gene |
end |
|
start |
||
Symbol |
The genes in the pathway |
|
Other columns |
Chrom |
|
Pathway |
The pathway(s) the given gene belongs to (pathway annotation is based on RACTOME, KEGG, WikiPathways, HGMD, Pathway_Interaction_Database, BIOCYC). |
|
set_orig_gene |
The genes in the given genelist |