Genes to paralogs¶
The Genes to paralogs dialog takes a list of genes and matches each gene to its corresponding paralogs. Alternatively, a list of paralogs can be created for all genes except those selected as input.
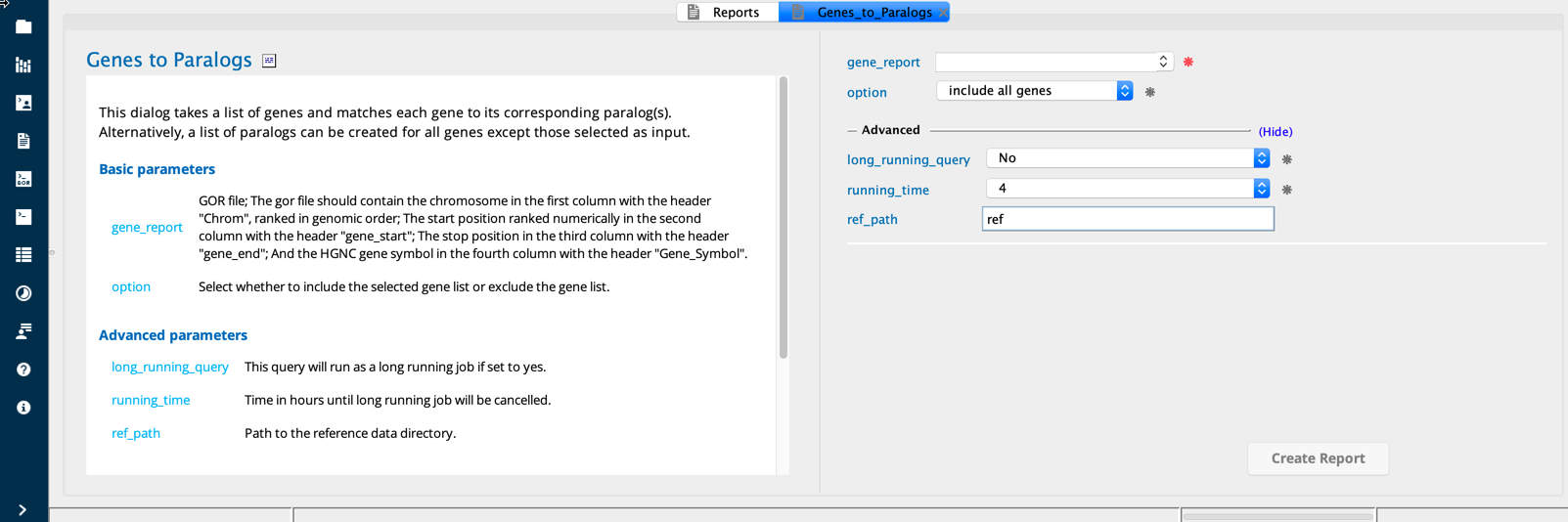
Genes To Paralogs module in Sequence Miner¶
Example use case¶
The user has a list of genes relevant to a phenotype and wishes to identify all the corresponding paralogs.
Description of the algorithm¶
Ensembl gene symbols, aliases, OMIM IDs, GO terms, and paralogs are stored in the ensgenes.map file in the ref folder in the Sequence Miner File Explorer. The unique genes in the Gene_Symbol column of the input gene report are mapped to matching Ensembl gene symbols and their corresponding Gene Paralogs listed in the ensgenes.map file.
Interpreting the output¶
A new column showing the list of paralogs is added and labeled “paralog_genes”.
Column descriptions¶
Group |
Column |
Description |
|---|---|---|
gene |
end |
|
start |
||
Symbol |
||
Other columns |
Chrom |
|
paralog_genes |
The paralogs of the given gene as defined by Ensembl |